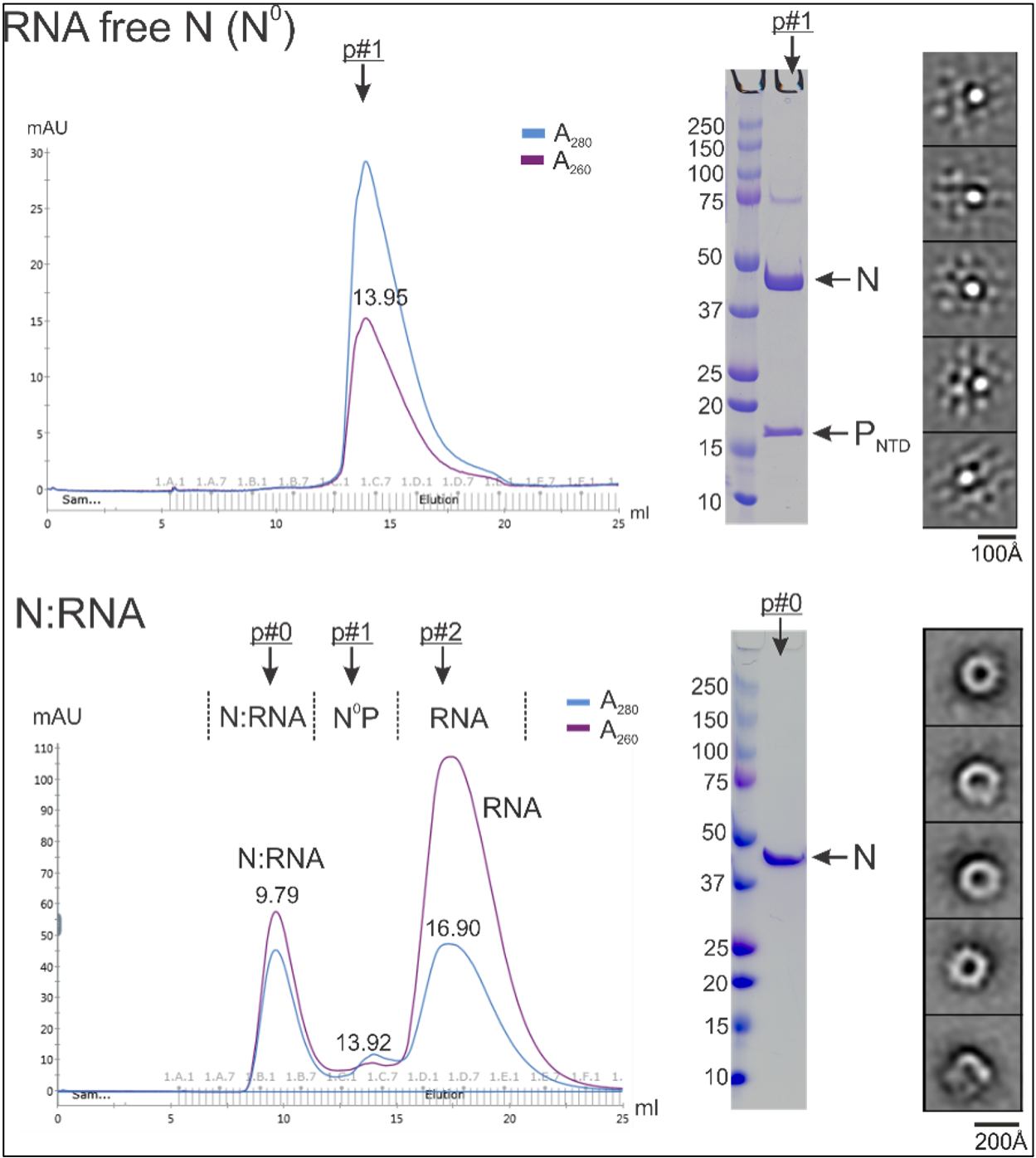
Lab Research and Projects
The Liang Laboratory focuses on integrating the power and advantages of different imaging system modalities, including those using electrons, x-rays, and visible lights, and scrutinizing how macromolecular machinery functions in various biological processes in real-time at unprecedented resolution. Capitalizing on breakthroughs in detector technology and image processing algorithms, the Liang laboratory is particularly interested in using cryo-electron microscopy (cryo-EM), the technology that won the 2017 Nobel Prize in Chemistry, as a primary tool to visualize macromolecular complexity. Cryo-EM is exceptionally well suited for molecular systems traditionally challenging for structural characterization, including membrane proteins and large and heterogeneous assemblies. Also, we employ X-ray crystallography and fluorescence light microscopy to provide complementary structural and dynamic information for challenging systems of significant biological importance.
Primary Projects
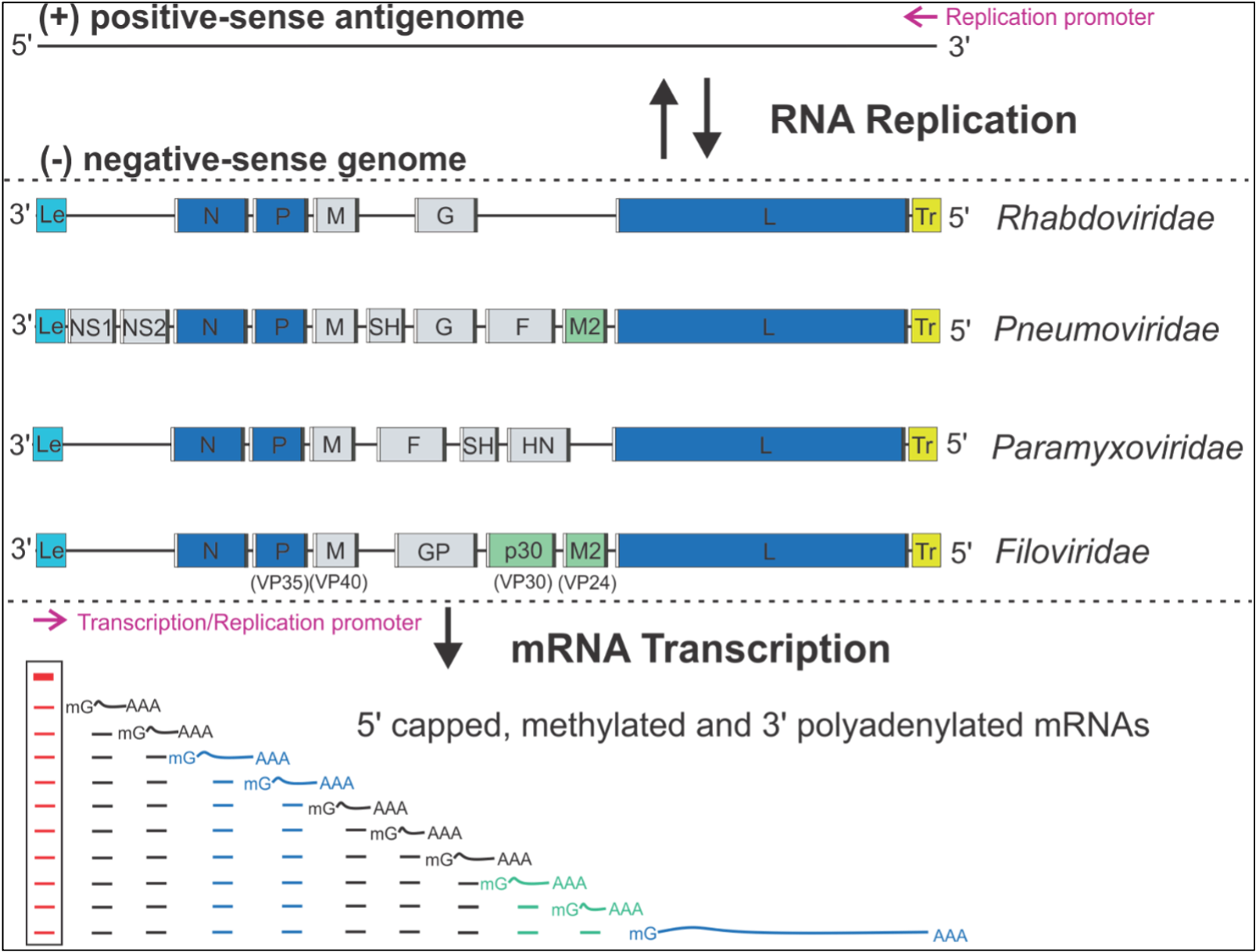
One such challenging system is the RNA synthesis machinery of a class of pathogenic and sometimes deadly non-segmented negative-sense (NNS) RNA viruses, including measles (MV), rabies (RABV), Ebola (EBOV), Marburg (MARV), and respiratory syncytial virus (RSV). RNA synthesis is central to the life cycle of NNS RNA viruses, carried out by a multi-functional RNA polymerase. The structural basis of the RNA synthesis machinery remains mostly unclear.
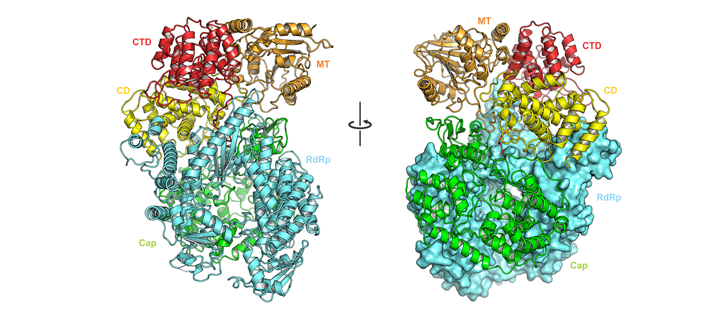
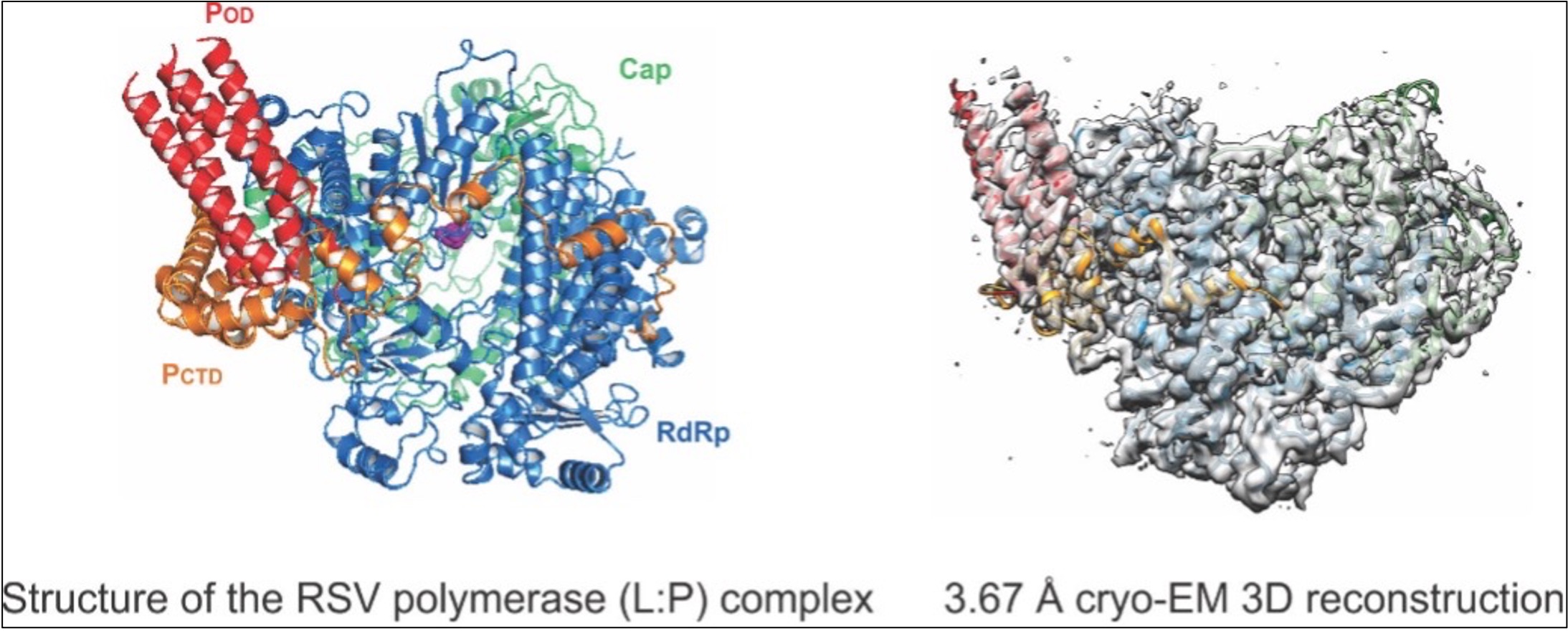
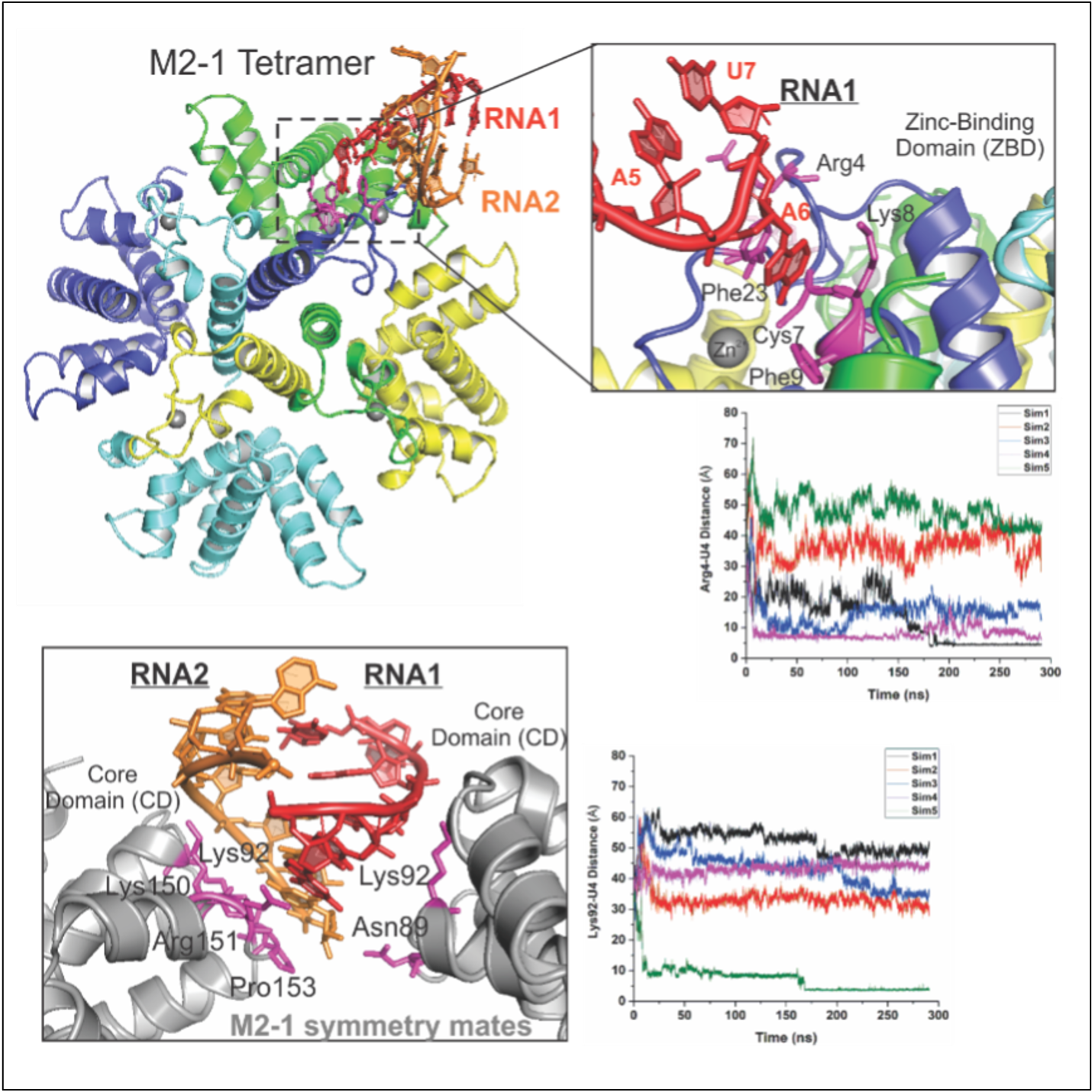
The Liang laboratory is devoted to understanding the structure and mechanism of the RNA synthesis machinery of RSV, the leading cause of severe respiratory tract diseases in young children and older adults, and immunocompromised in the United States and worldwide. RNA synthesis by the RdRP of RSV is essential for viral pathogenesis. The Liang laboratory aims to establish an RNA synthesis platform for RSV, elucidate how this RNA synthesis machine functions, and identify potential antiviral therapeutic targets for more effective treatment. Our immediate research goal would be to decipher the molecular architecture of the RNA synthesis machinery of these viruses using cryo-EM and x-ray crystallography. Our research could lead to the rational design of effective antiviral drugs to block RSV activity. Such drugs would substantially reduce serious RSV infections in humans and greatly improve human health.